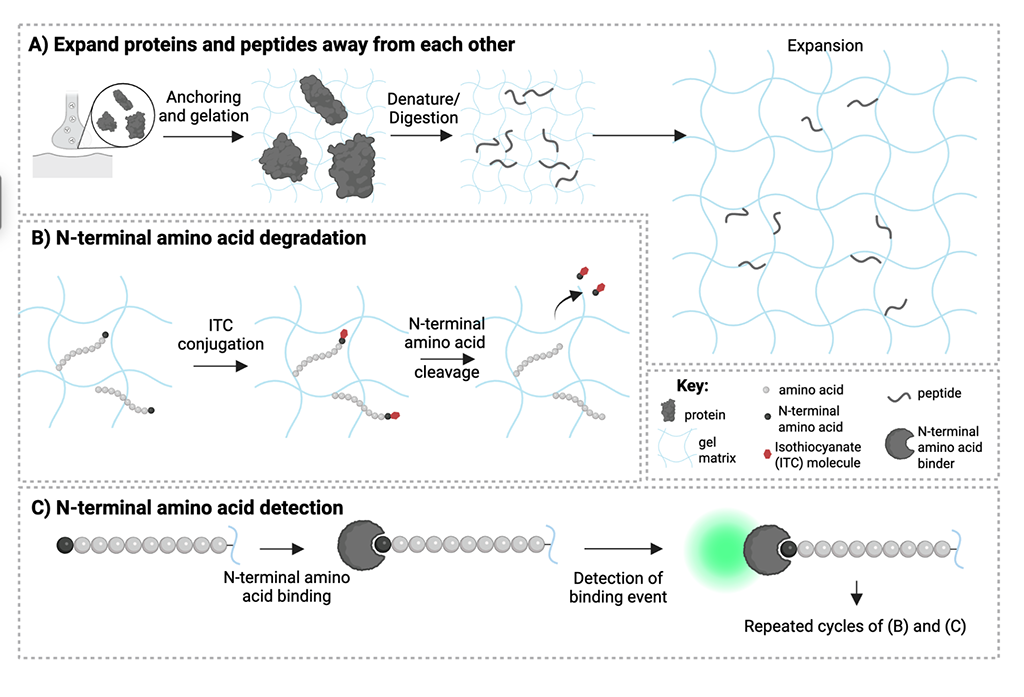
The ability to map protein identity, with resolution sufficient to infer interactions, would support analysis of how proteins work together, or malfunction, in biological processes and diseases. Although several emerging technologies aim towards single-molecule protein sequencing, they require proteins to be removed from the nanoscale spatial context of cells and tissues. Expansion microscopy (ExM) has facilitated a diversity of chemical analyses by isotropically separating molecules throughout a specimen after permeation via a charged hydrogel, followed by gel swelling. Here, we adapt key protein sequencing steps – Edman degradation and amino acid recognition – to the ExM gel context. Using testbed peptides in ExM gels, we show that N-terminal amino acids can be recognized over multiple cycles of in-gel Edman degradation. These results establish principles of in situ protein sequencing and provide a framework for future in situ protein sequencing developments, including the development of higher specificity and affinity amino acid binders.