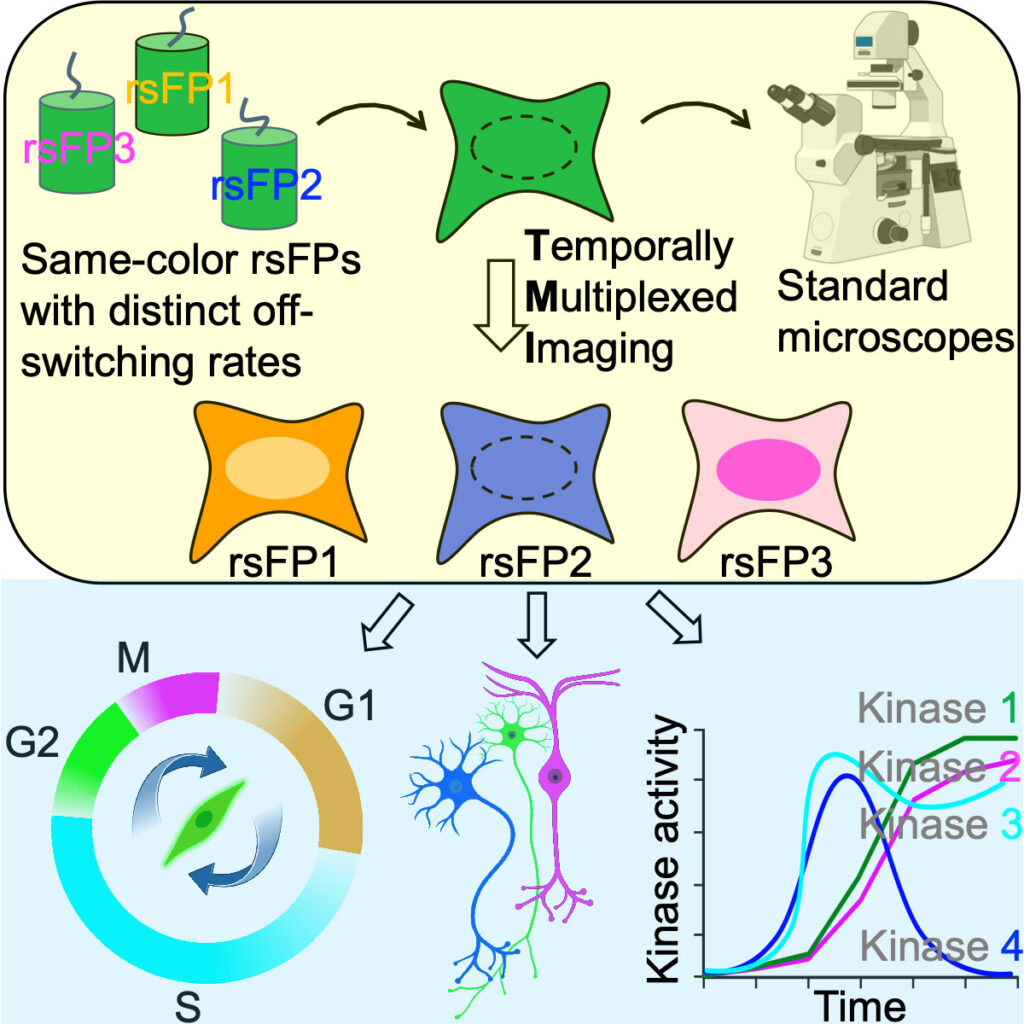
Molecular signals interact in networks to mediate biological processes. To analyze these networks, it would be useful to image many signals at once, in the same living cell, using standard microscopes and genetically encoded fluorescent reporters. Here, we report temporally multiplexed imaging (TMI), which uses genetically encoded fluorescent proteins with different clocklike properties—such as reversibly photoswitchable fluorescent proteins with different switching kinetics—to represent different cellular signals. We linearly decompose a brief (few-second-long) trace of the fluorescence fluctuations, at each point in a cell, into a weighted sum of the traces exhibited by each fluorophore expressed in the cell. The weights then represent the signal amplitudes. We use TMI to analyze relationships between different kinase activities in individual cells, as well as between different cell-cycle signals, pointing toward broad utility throughout biology in the analysis of signal transduction cascades in living systems.
Original biorxiv preprint: version 1 posted on August 22, 2022, and available [here]
