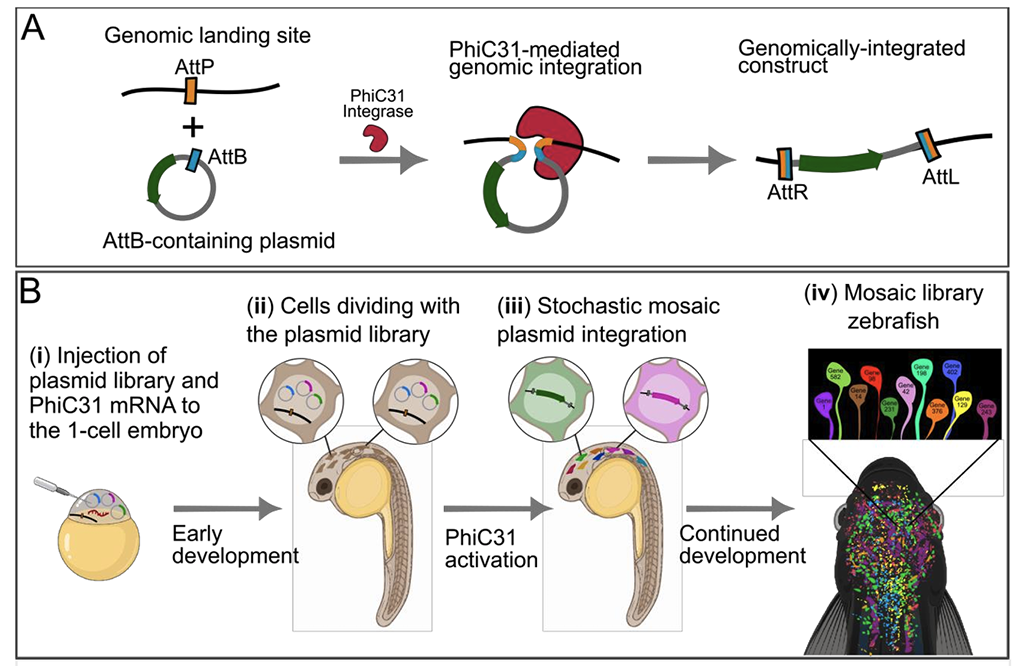
Functional screening through systematic deletion, editing or addition of libraries of genes is a powerful approach for discovering gene functions and developing improved molecular tools. However, due to the need for high throughput, such campaigns are typically conducted in vitro, leading to many discoveries, especially tools and therapeutics, which fail to translate in vivo. Tissue context, cellular physiology, and systemic regulation shape both tool performance and gene function in ways that simplified culture systems cannot predict. Pooled in vivo screening methods have the potential to enable screening within living animals while preserving the physiological context, but current approaches using viral vectors face three critical limitations: multi-transgene insertions per cell confound genotype-phenotype association, viral packaging constrains transgene size, and cell-type tropism restricts and biases targeting. Here, we introduce a zebrafish library transgenesis method that overcomes these limitations through delayed site-specific mosaic integration. We exploit a temporal delay between library microinjection with PhiC31 mRNA, and library integration, to allows the library to spread episomally throughout the developing embryo before integration begins. This produces mosaic animals where each cell independently integrates one randomly-selected library member, enforced by a single genomic AttP landing site. We demonstrate delivery of multi-kilobase transgenes with high library coverage of 1,378-1,989 unique integrants per animal, and single-transgene-per-cell in ∼99% of brain cells. This method provides a platform for direct in vivo screening of large transgene libraries with single-transgene precision, with potential applications in both biological discovery and tool development.
